How do we do it?
The technology behind PharmAI’s DiscoveryEngine allows for an unprecedented level of detail in the representation of drug-target complexes. Besides geometric comparison of binding sites based on local alignments, non-covalent interactions were included as they significantly improve the prediction of binding site similarities. Over many years, PharmAI refined and improved this technology and applied it to reposition known drugs to new targets for cancer or malaria. At its core, pharmAI’s technology leverages 3D structure of drug-target complex, calculates similarities to produce predictions, and ranks them using artificial intelligence.
Interactions Unique as Your Fingerprint
PharmAI's DiscoveryEngine translates protein-ligand complexes into unique fingerprints. This allows us to quickly compute similarities across arbitrary chemical libraries in order to predict new compounds. We can even use the fingerprint for the fingerprint even for the prediction of potential off-targets.
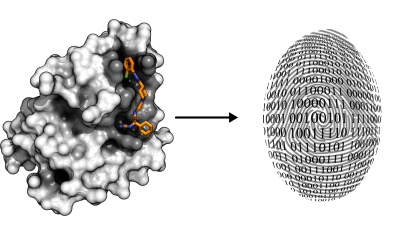
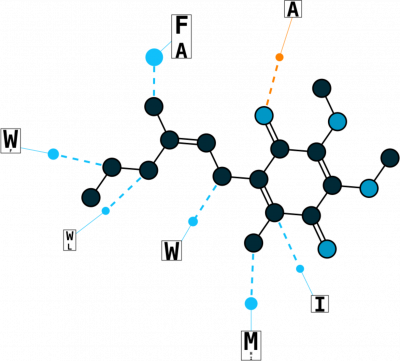
Analyzing Binding Sites
The key to pharmAI's disrupting technology is the exploitation of geometric binding site properties and non-covalent interaction patterns. The unique combination of these two approaches allows us to generalize ligand binding and predict new scaffolds.
Our Steps to Find New Scaffolds and Off-Targets
Binding Site Analysis
Interaction Analysis
Fingerprinting
Superior Performance
Hit rate 500x higher than HTS rates, 8x-3x improvement over than state-of-the-art methods.
Comparison to Industry Leaders
Our hit rate outperforms methods used by industry leaders:

